POSTS
Hit and miss for sloppy transcription factors
For many years, we have been trying to understand how LacI and other transcription factors (TFs) scan the DNA in search of their specific binding sites; with the help of simulations and indirect methods, we have arrived at a detailed model of the search mechanism. However, we have not been able to directly monitor the process due to shortcomings in the available methods. It is not without pride that we now announce these limitations to be history.
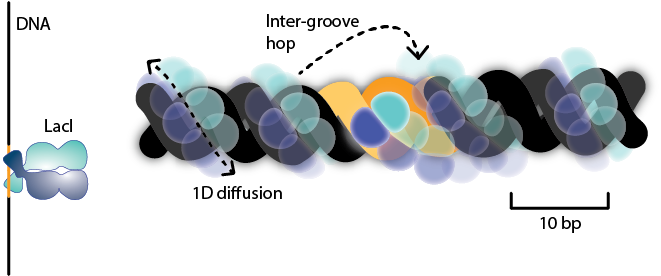
In a study that will shortly be published in Nature, we follow the movement of LacI sliding on DNA in real-time, and conclude that although the TF moves around the DNA in a spiral motion, it does not faithfully track the major groove. We combine the real-time imaging with a smFRET study which corroborates these findings. In summary, we show how the TF is trading accuracy for speed when scanning the DNA. Although the TF will frequently miss the target site, it will revisit the sequence enough times to ensure rapid binding.
Read the whole story in Nature.