POSTS
Single Molecule Kinetics in Living Cells - A Review
In the past decades, advances in microscopy have made it possible to study the dynamics of individual biomolecules in vitro and resolve intramolecular kinetics that would otherwise be hidden in ensemble averages. More recently, single-molecule methods have been used to image, localize and track individually labeled macromolecules in the cytoplasm of living cells, allowing investigations of intermolecular kinetics under physiologically relevant conditions. In a recent review, accepted for publication in Annual Review of Biochemistry, we illuminate the particular advantages of single-molecule techniques when studying kinetics in living cells and discuss solutions to specific challenges associated with these methods.
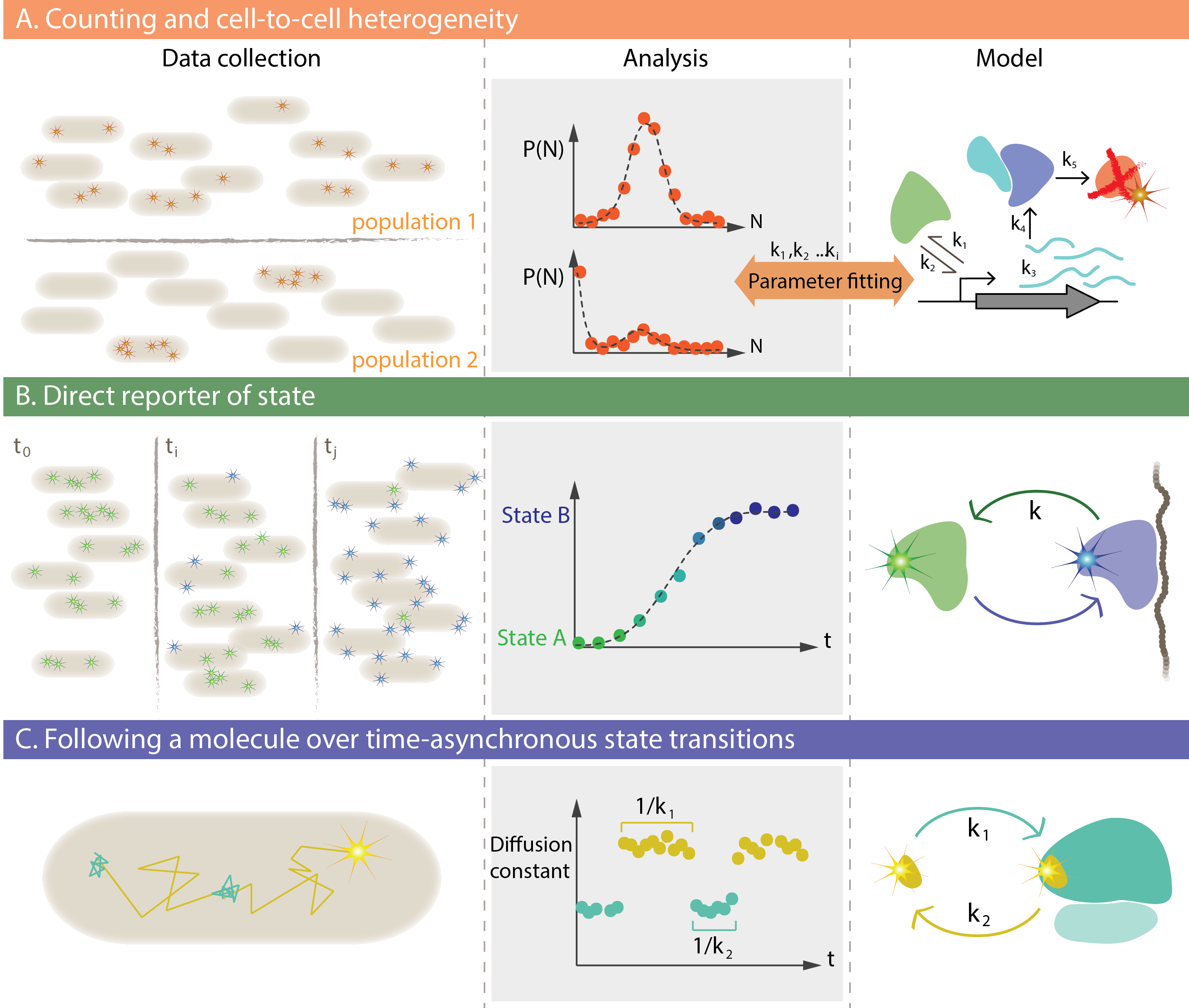
The image describes three general use cases for dynamic single molecule methods in living cells. (A) To determine the number of a particular molecular species and how this number varies over time, in space or between cells. (B) To measure the fractions of mobile and immobile molecules, how these fractions are distributed in the cell and whether a particular binding site is occupied or not. © To determine transition rates by monitoring state changes one molecule at a time.