POSTS
New analysis pipeline put to the test
This study was recently published in Nature Chemical Biology. In short, we track the movement of individual intracellular [Cy5]tRNAs by stroboscopic laser illumination with 1.5 ms laser illumination per 5 ms camera exposure frame. Combining symmetry-based spot detection and maximum a posteriori estimates of position and localization uncertainty with the u-track algorithm, we follow the movement of single molecules up to more than 100 frames. The apparent diffusion is estimated from mean-squared displacement (MSD) plots of trajectory segments.
To quantify binding events in terms of the accompanying changes in diffusion constant, we use a hidden Markov model (HMM) approach, which models the trajectories as random (Markovian) transitions between a set of discrete (hidden) states with different diffusion constants, accounts for point-wise localization errors and motion blur, and learns the underlying sequence of state transitions from data. The HMM algorithm performs maximum-likelihood inference of model parameters, and we use Akaike’s information criterion (AIC) to determine the appropriate number of diffusive states to fit the data.
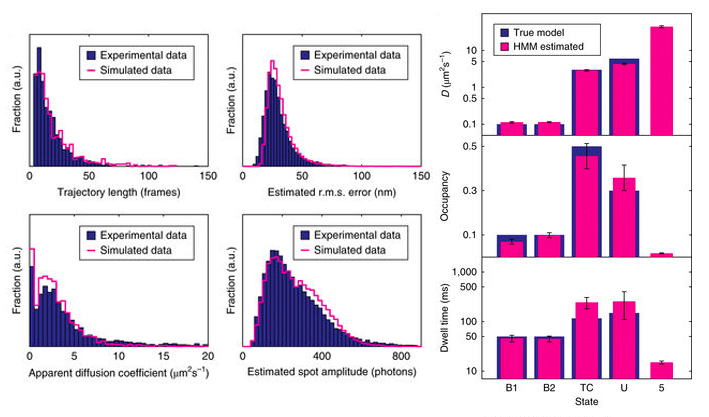
The analysis pipeline was validated by synthetic data based on simulated trajectories, camera noise and point spread functions. The statistics generated from the simulated data was very similar to the statistics obtained from the experimental movies and the analysis faithfully recovers the parameters of the true model (see image below). We are thus confident that the analysis pipeline is capable of producing reliable results.
The Cy5-labeled tRNAs display diffusion patterns consistent with what would be expected for a tRNA being used repeatedly by the protein synthesis machinery, i.e., random fast diffusion interrupted by immobilization events, possibly reporting on binding of the tRNAs to an mRNA-tethered ribosome.
The study is commented by Achillefs Kapanidis and Mathew Starcy in their News and Views article Tracking tRNA packages.