POSTS
Cell cycle regulation unveiled
How the cell cycle is regulated in bacteria has been examined and debated since the first model was put forth by Cooper and Helmstetter in the 1960s. Using a combination of microfluidics and image analysis of E. coli with fluorescent labels on the replication machinery, we have been able to study how the replication and division cycles are coupled in individual cells. This made it possible sort out what is the source if variability in cell sizes and generation times. In brief, all cells initiate at a constant volume per chromosome. The time from initiation to division is given by the individual cells growth rate which is variable and is the major cause of the variation in cell sizes. It has been a long project, 5 years, but it turned out very nicely in the end.
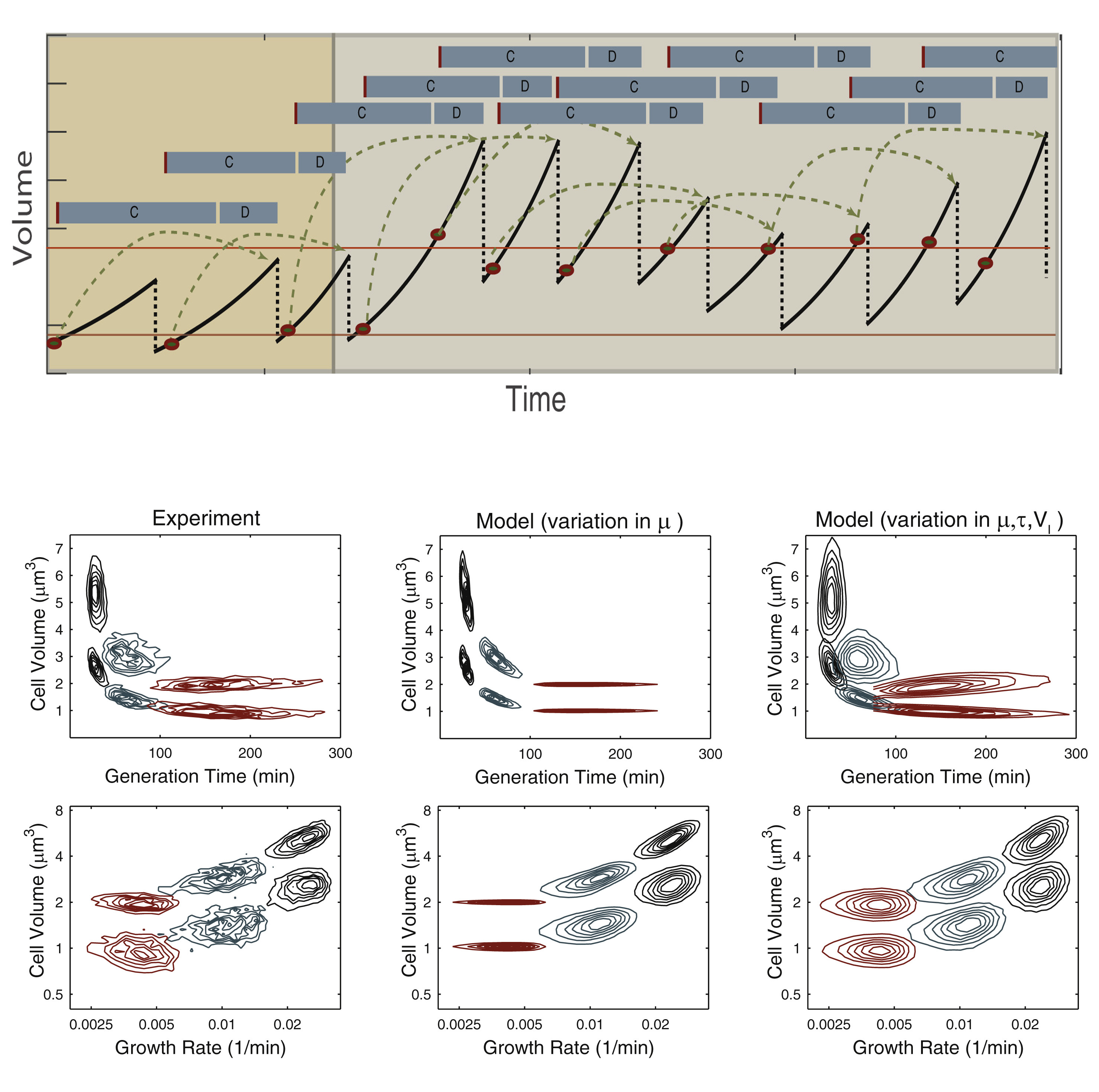
(Top) The cell volume (black solid line) of a single cell lineage simulated according to the single-cell CH model throughout an upshift in growth conditions. The growth rates are sampled from the observed distributions. When the cell passes the initiation volume(s) (red lines), a division event is triggered (green arrow) after a time period corresponding to the C and D periods (gray bars). Variation can be introduced in growth rates, initiation volume, or C+D period. (Bottom) Demonstration of how the model performance is improved as cell-to-cell variation is introduced in growth rate (mu) as well as in initiation volume (V) and C+D period (tao).
Check out the publication in Cell